Reading time: 4 minutes
Deepika Jayaprakash
What is common to eukaryotic organisms like animals, plants, fungi, and unicellular organisms?
Ubiquitous Ubiquitin!
Ubiquitin (ub) is a small protein found in all eukaryotes that when attached to its target protein can bring about a myriad of cellular processes. The first observed effect of ubiquitination was degradation of the target proteins by a protease enzyme complex called proteasome, leading to regulation of the respective cellular pathways. The ground-breaking discovery of the ubiquitination cascade (Fig 1) in early 1980s by Hershko, Rose, and coworkers led the 3 scientists to receive the Nobel prize in Chemistry in the year 2004. Following the discovery, the non-degradative consequences of ubiquitination such as DNA damage repair, cell cycle regulation, signaling to control inflammation and more were identified. The fate of ubiquitination depends on the pattern (Fig 2) and site of modification that occurs on target proteins.
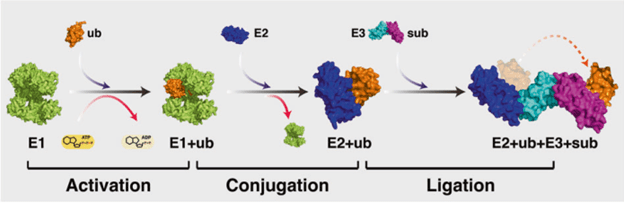
Fig 1: Ubiquitination cascade – the target protein is covalently bound with the ubiquitin molecule through a series of reactions starting with activation of ubiquitin by activating enzyme E1, followed by transfer of ub to E2 conjugating enzyme, and finally ub is bound to the target protein/substrate(sub) by the E3 ligase. Image source
Since ubiquitination of proteins are involved in a gamut of biological processes, any mutation in the ub system leads to several disorders, including cancer. There are about 600 genes coding for different E3 ligases in the human body, the highest in comparison to other components of the ub system. Based on the structure, E3s can be classified into three main subfamilies:
– RING (really interesting new gene) finger domain-containing E3s
– catalytic HECT (homologous to E6-AP C-terminal) domain containing E3s
– the RING-in-between-RING (RBR) E3s
How do E3 ligases support cancer growth?
Genetic variations/mutations of E3 ligases have been reported in many cancers. Mutated E3s have the ability to degrade tumor suppressor genes. Tumor suppressor genes, as the name suggests, are genes that keep a check on cell division thereby ensuring regulation of the cell cycle. Consequently, when the suppressor gene is lost, it leads to uncontrolled cell division that may lead to cancer and help the already formed cancer to grow bigger. For example, the well-known tumor suppressor gene p53, otherwise called guardian of the genome, is regulated by E3 ligase MDM2. MDM2 ubiquitinates p53 thereby resulting in its degradation and subsequent uncontrolled cell growth. MDM2 is overexpressed in several cancers leading to unchecked cell growth.
On the other hand, mutations in certain E3 ligases that regulate oncogenes (genes that increase susceptibility to cancer) increase the levels of oncogenes thereby aiding the cancer cells to grow faster.
Additionally, alterations in certain other E3 ligases called IAPs/Inhibitors of apoptosis-related proteins can evade apoptosis, the programmed mechanism to kill the damaged cells in the body, thereby increasing the numbers of damaged cells. Furthermore, E3 ligases promote metastasis, where cancer cells travel through blood or lymph to lodge to a distant site, by Epithelial mesenchymal transition (EMT). EMT is a process where the epithelial cells gain mesenchymal cell properties i.e. lose tight cell-cell adhesions, become migratory and invasive; thereby detaching and metastasizing to a different location.
Exploiting E3 ligases to treat cancer
Since E3 ligases are involved in several important cellular processes during development and homeostasis, they were generally considered not good candidates to target. However subsequently, E3 inhibitors with 3 different modes of action (Fig 3) were developed:
– small molecule inhibitors that disrupt binding substrate with E3 ligases
– engineered E3 variant to inhibit the ligase
– molecules which change the conformation of the protein directing them to be degraded
Chemical reactivation of ubiquitination can be achieved by proteolysis targeting chimeras (PROTACs) and molecular glues (Fig 3) used for targeted protein degradation. PROTACs are bifunctional small molecules with a specific E3-binding moiety, a targeted protein-binding moiety and a connecting linker. PROTACs artificially induce ubiquitination and subsequent degradation of the targeted protein. Molecular glues work in a similar manner to PROTACs, but due to absence of the chemical linker, the physical proximity between the ubiquitination and the subsequent degradation is not achieved.
Indirect strategies like inhibition of proteasomes results in the induction of cell cycle arrest and apoptosis via modulation of several pathways including stabilization of p53, and activation of apoptosis. Up to date, three proteasome inhibitors namely bortezomib (2003), carfilzomib (2012) and ixazomib (2015) have been approved by the US Food and Drug Administration (FDA) for the treatment of patients with relapsed and/or refractory Multiple Myeloma.
With several scientists working worldwide on E3 ligases in cancer, I am certain that we will soon have more definitive therapies exploiting the E3s in the near future to combat the disease progression.
Edited by Aya Elmeligy
Leave a comment