Reading time: 7 minutes
Nayela Chowdhury
Conventional 2D and 3D models of cancer have widely been utilized in pre-clinical drug development; however, neither of the platforms mimics the dynamic interactions that occur in living tumors. In vivo studies (for example, in small animal models) have been central in translational cancer research and have contributed largely to understanding disease progression and development of new treatments. The last decade has seen an increase in the use of patient-derived xenografts (PDX) and genetically engineered mouse (GEM) models over long-established cell lines in preclinical drug screening. PDX models incorporate patient tumor cells into immunodeficient mice to resemble the original tumor in the patient. Therefore, PDX models can be personalized for individual patients, which is invaluable in studying drug response and drug resistance over a diverse population. At the same time, PDX models generate stable tumors which preserve gene expression, mutation, histopathology, and inflammation of the patient tumor [1]. However, PDX models are immunocompromised, utilizes rodent or host stroma, and lack standardization [2]. In addition, not all types of tumors can be engrafted in PDX models [3]. On the other hand, GEM models are mouse models that are genetically engineered to carry the same mutations in cancer development. One or more of their genes can be manipulated to express, overexpress, or delete a gene involved in the pathophysiology of a corresponding human tumor [4]. Contrary to PDX models, GEM models employ a functional immune system to study drug response, kinetics, and tumor metastasis. Likewise, they also maintain tumor stroma and vasculature [5]. However, these models are associated with high costs and more importantly, GEMs do not simulate human disease states as these tumors are strictly rodent tumors [6]. GEM models cannot be personalized and therefore, have no utility in studying therapeutics over a diverse human population. Furthermore, aside from regular ethical concerns, in vivo studies in animals such as mice [7] and non-human primates [8, 9] have not only failed to replicate equivalent therapeutic response in humans but also failed to accurately predict fatal adverse effects.
Tumor-on-a-chip for Drug Discovery
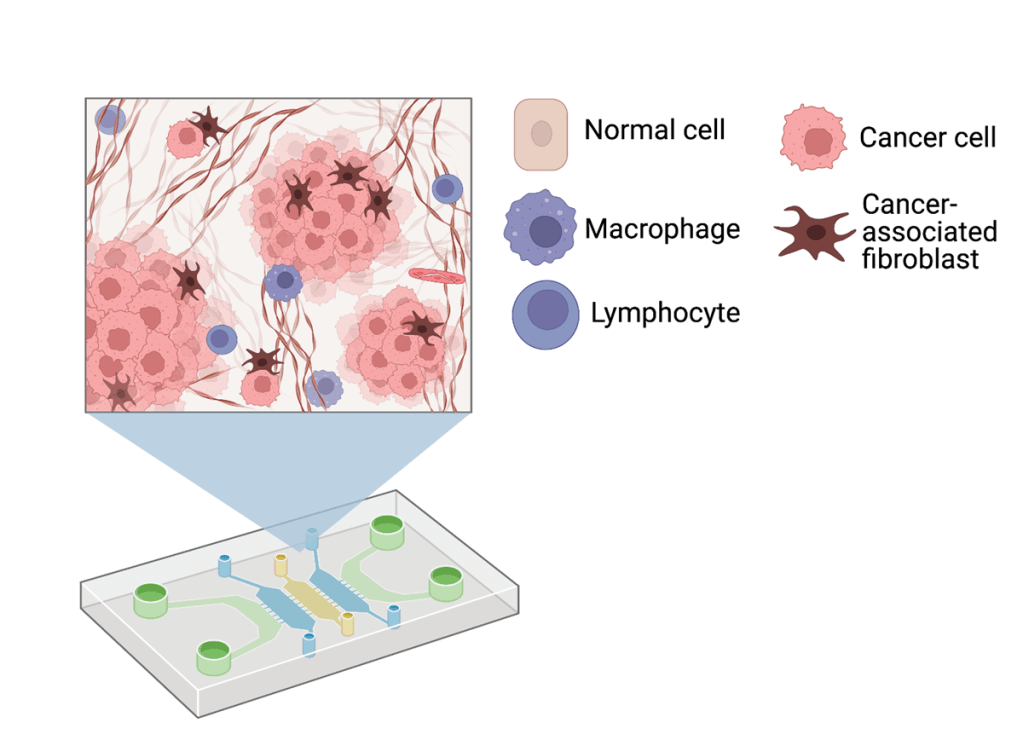
Microfluidics-based tumor-on-a-chip (TOC) systems provide a promising approach to address these challenges. Microfluidics is a branch of science and technology that manipulates and studies dynamic behavior of miniscule amounts of fluid in micrometer sized channels. These chips can house human cells along with their extracellular matrix (ECM) to exhibit organ-level functions and differentiation, which can be observed in real time (Figure 1). The ECM is crucial to cancer cell proliferation and is involved in cell adhesion, invasion, and migration to secondary sites. Since these chips can incorporate human tissues, they provide a platform that is biologically relevant without raising ethical dilemmas.
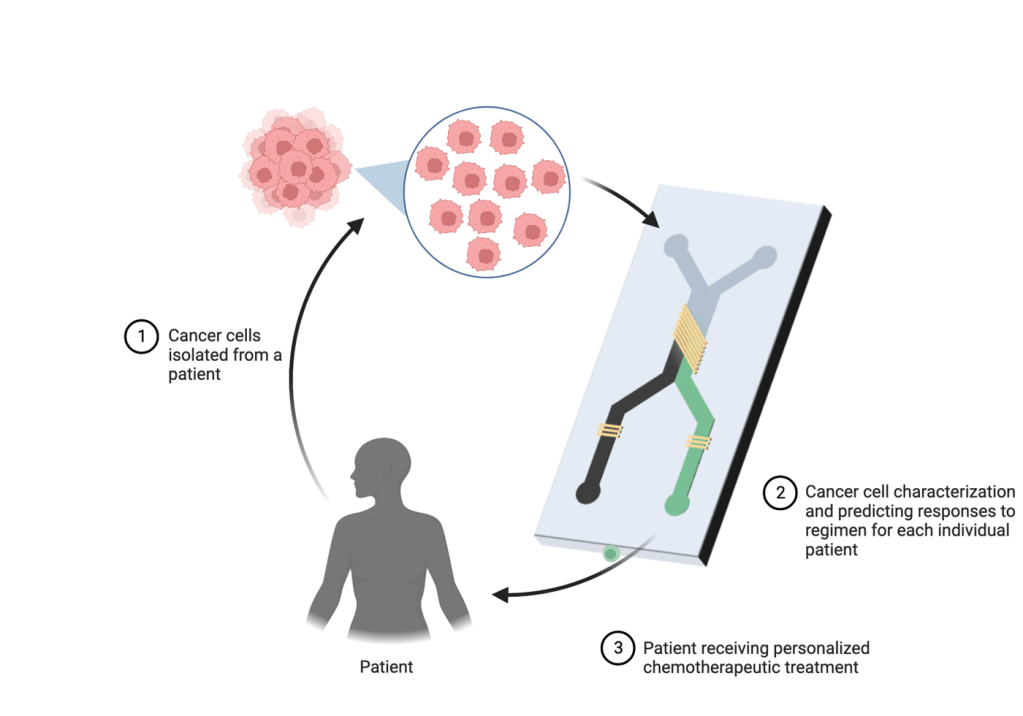
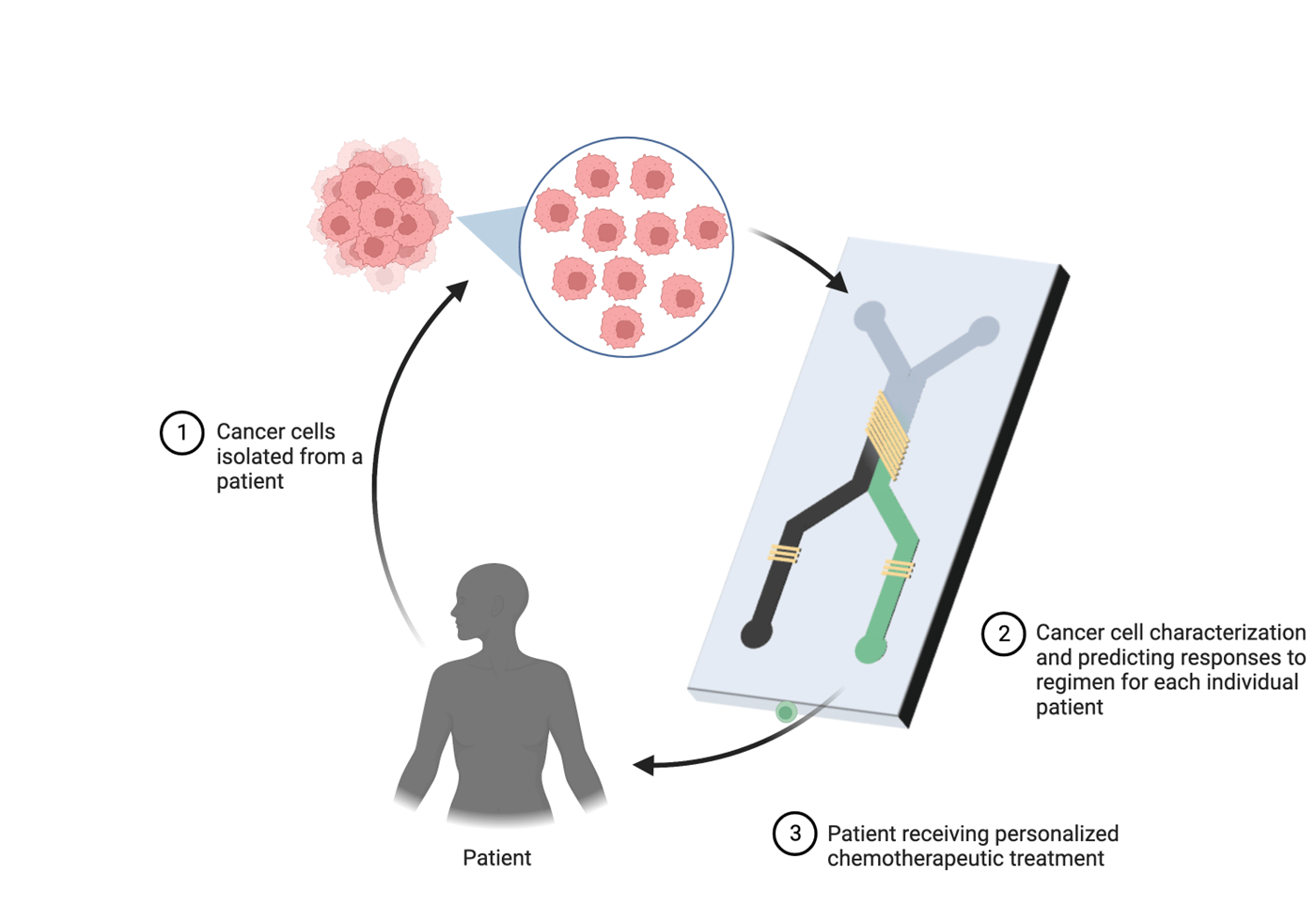
The basic elements of microfluidic chips consist of valves, microchannels and chambers that are interconnected for cell culture and precise control of fluid flow that create a dynamic environment for the cells to grow. Moreover, fabrication of microfluidic chips is relatively inexpensive and reduces the cost of the drug development process in the long term. Microfluidic devices are also diverse in terms of utility, and have been employed in studying personalized medicine, drug delivery, tissue engineering and point-of-care-testing. For instance, Xu et. al. [10] integrated monocultures and cocultures of primary lung cancer cell lines and stromal cells in one microfluidic device on separate interconnected chips. These cells were treated with chemotherapeutic agents such as gefitinib, paclitaxel, cisplatin and gemcitabine (individually and in combination) and results were applied to fresh lung tissues biopsied from eight patients in different stages of cancer. In all cases, effective response to single or combination therapy was different for each patient, implying that the chip can be useful in providing guidance for personalized therapy for cancer. This implicates the utility of these chips in identifying the best drug regimen for individual patients to maximize response to chemotherapy (Figure 2). Choi et. al. [11] administered paclitaxel, an anti-cancer agent commonly prescribed in breast cancer as a first line treatment, in ductal carcinoma spheroids on a chip. Ductal carcinoma was recapitulated using cocultures of mammary epithelial cells and primary mammary fibroblasts. Remarkably, epithelial cells alone experienced negligible cytotoxicity but paclitaxel effectively inhibited growth of the ductal carcinoma spheroids compared to untreated tumor spheroids. Similarly, Gampala et. al. studied the effects of single and combination treatment in an interstitial tumor-microenvironment-on-chip (iT-MOC) containing cocultures of pancreatic cancer cells and associated fibroblasts. Dynamic blood-borne drug transport was mimicked using a capillary vessel, and it showed that combination treatment of Redox factor 1 (Ref-1) inhibitor and Devimistat (TCA cycle inhibitor) showed significant specificity in inhibition of pancreatic cancer cells proliferation while preserving the fibroblasts [12].
Applications of TOC
TOC technology aims to mimic in vivo tissue environment in vitro on a chip. In addition to solid tumors, other models recapitulating biological processes in cancer development such as angiogenesis, invasion, and metastasis have been developed over time to mimic cancer niche in patients. Sung et. al. [13] studied the pathophysiology of ductal carcinoma from localized to invasive state in a compartmentalized model housing mammary epithelial cells spatially cultured with and without human mammary fibroblasts. Remarkably, mammary epithelial cells acquired rapid invasive potential on contact with the fibroblasts in contrast to monocultures, providing valuable information on cell signaling and potential therapeutic targets to inhibit invasion. Likewise, metastasis has also been replicated on a microfluidic system to capture the microenvironment that supports growth and development of metastases of triple negative breast cancer into the bone [14]. On another note, angiogenesis is the development of vascular networks within a tumor that provides with nutrients crucial for its rapid expansion and growth. This biological process has already been modeled on a chip by Nguyen et. al. [15] who seeded endothelial cells in a collagen channel and exposed to several angiogenesis inducing factors. Interestingly, under those conditions the cells differentiated to form neo vessels between the channels resembling vascular networks in vivo. Using this model, a cocktail of angiogenic factors that induces the highest angiogenic response was identified.
TOC: Challenges and Perspectives
Tumor on a chip technology is still in its infancy and is faced with several engineering challenges. Individual organs and multi-organ systems imitated on chips will require standardization and reproducibility to replace traditional cell based analytical platforms in drug discovery. Future improvements may include increasing tissue longevity and integration of biosensors to monitor pH, temperature, oxygen levels, glucose and toxin buildup while reducing complexity of the chip for high throughput analysis. Once regulated, organ-on-a-chip platforms can potentially become the new standard of biological models required in different stages of drug development to reduce the financial burden in each phase. Consequently, they can increase success of clinical trials by identifying drugs with unsatisfactory therapeutic and pharmacokinetic profiles in early stages of drug discovery. Continued advances in optimization of structure, fabrication techniques, types of biomaterials employed, and cell culture methods can essentially diminish attrition of drugs and facilitate expedited discovery of new drugs in pipeline.
Edited by Ana Isabel Castillo Orozco
Works Discussed
1. Kuracha, M.R., et al., Patient‐derived xenograft mouse models of pseudomyxoma peritonei recapitulate the human inflammatory tumor microenvironment. Cancer medicine, 2016. 5(4): p. 711-719.
2. Gao, H., et al., High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nature medicine, 2015. 21(11): p. 1318.
3. Hidalgo, M., et al., Patient-derived xenograft models: an emerging platform for translational cancer research. Cancer discovery, 2014. 4(9): p. 998-1013.
4. Richmond, A. and Y. Su, Mouse xenograft models vs GEM models for human cancer therapeutics. 2008, The Company of Biologists Ltd.
5. Gopinathan, A. and D.A. Tuveson, The use of GEM models for experimental cancer therapeutics. 2008, The Company of Biologists Ltd.
6. McMillin, D.W., J.M. Negri, and C.S. Mitsiades, The role of tumour–stromal interactions in modifying drug response: challenges and opportunities. Nature reviews Drug discovery, 2013. 12(3): p. 217.
7. Seok, J., et al., Genomic responses in mouse models poorly mimic human inflammatory diseases. Proceedings of the National Academy of Sciences, 2013. 110(9): p. 3507-3512.
8. Watkins, D.I., et al., Nonhuman primate models and the failure of the Merck HIV-1 vaccine in humans. Nature medicine, 2008. 14(6): p. 617.
9. Bailey, J., An assessment of the role of chimpanzees in AIDS vaccine research. 2008.
10. Xu, Z., et al., Application of a microfluidic chip-based 3D co-culture to test drug sensitivity for individualized treatment of lung cancer. Biomaterials, 2013. 34(16): p. 4109-4117.
11. Choi, Y., et al., A microengineered pathophysiological model of early-stage breast cancer. Lab on a Chip, 2015. 15(16): p. 3350-3357.
12. Gampala, S., et al., Ref-1 redox activity alters cancer cell metabolism in pancreatic cancer: exploiting this novel finding as a potential target. Journal of Experimental & Clinical Cancer Research, 2021. 40(1): p. 1-24.
13. Sung, K.E., et al., Transition to invasion in breast cancer: a microfluidic in vitro model enables examination of spatial and temporal effects. Integrative Biology, 2011. 3(4): p. 439-450.
14. Bersini, S., et al., A microfluidic 3D in vitro model for specificity of breast cancer metastasis to bone. Biomaterials, 2014. 35(8): p. 2454-2461.
15. Nguyen, D.-H.T., et al., Biomimetic model to reconstitute angiogenic sprouting morphogenesis in vitro. Proceedings of the National Academy of Sciences, 2013. 110(17): p. 6712-6717.
Leave a comment