Reading time: 5 minutes
Kaye Alcedo
Even before President Nixon’s declaration of the “war on cancer” in 1971, the journey towards a cure was nothing but a rollercoaster ride. Many promising cancer drugs are tested in human clinical trials but ultimately fail primarily because they demonstrate inadequate efficacy or safety, costing billions of dollars, increasing the burden on patients, and prolonging the search for a cure. One hurdle that Dr. Don Ingber, from the Wyss Institute, described during his recent talk at UNC-Chapel Hill was how the current experimental models may not be good enough at predicting the efficacy and safety of a drug in patients.
In a previous post here at Oncobites, Manisit discussed the process of drug development, which culminates in human clinical trials. However, before a drug gets to that stage in the pipeline, it is tested through preclinical studies using two models: cell-based and animal-based. Drugs are almost always (with a few exceptions) initially tested on a monolayer of cells plated on a petri dish to determine their therapeutic effects. Subsequently, they are tested in animal models to determine their efficacy and safety before studies in humans. The cell-based model is advantageous in that it is simple. It allows for identification of important molecules within a particular cell that can be targeted by a drug. These cells can be from humans or animals; they can be normal or cancerous cells. This model enables scientists to test how a certain drug can change how cancer cells behave, look, grow, and/or survive. However, the simplicity of this model is also a disadvantage because, in reality, cancer cells grow in 3-dimensional structures, and are part of a “microenvironment” that is comprised of other types of cells and molecules. Thus, the effects of a drug on cells in a petri dish may not be to the same extent as when other types of cells are present. Further, many complex processes such as absorption, metabolism, and excretion can significantly change how drugs behave in the body, compared to cells on a petri dish. In order to overcome these limitations, drugs are tested in animal models.
As mentioned in the previous post, animal models enable the study of how “drugs function and interact with . . . complex biology.” Drugs are typically taken orally or given by intravenous (IV), skin, or muscle injections. Once the drug gets in the body, it may have to be transported to the liver to be processed and become activated. Subsequently, the active drug often has to reach a specific target in order for the therapeutic effects to occur. The journey of the drug can be long before it gets to the target. The drug has to pass through many different “microenvironments,” surviving harsh conditions in the stomach, passing through the cells in your gut to enter your body, getting filtered and degraded by enzymes in the liver, and interacting with a host of highly active proteins found in the blood. Last but not the least, in the case of cancer drugs, they must pass through the surrounding tissues that encase and protect the cancer cells. As you can see, the interaction of the drug is often not only with its specific target, but also with other cell types in the body, posing a risk of potential side effects. Thus, it is imperative to understand how the drug affects the target using the cell-based petri dish model, and also to study its effectiveness and safety using animal-based models. However, it is worth stating that animals are not humans; the “microenvironments” of the two have many differences that may account for why some drugs that have worked in animals have failed in human studies.
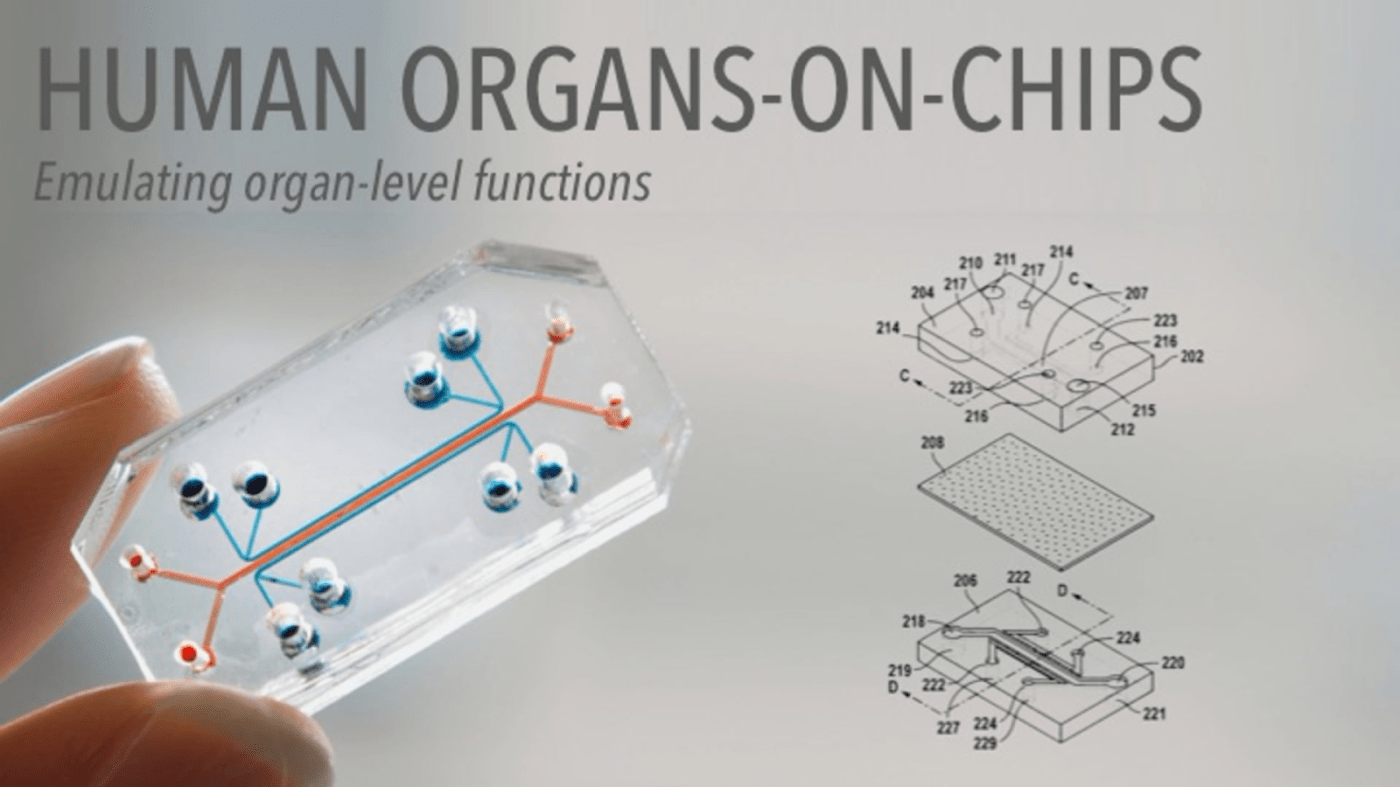
A promising alternative approach to circumvent the limitations of cell-based and animal-based models is the human-organs-on-chips technology that Dr. Ingber and his team pioneered. Their aim was to recreate complex, physiologically functioning organs without the need for animals studies. They were able to achieve this by incorporating multiple types of human cells and “placing them in a dynamic and mechanically relevant organ-specific microenvironment” in small, microchip-like devices. For example, they cultured human lung cells (epithelium) on top of a thin, porous, and flexible membrane, and human blood vessel cells (endothelium) at the bottom (Fig A). The top part of the device contains air and the bottom part contains fluid, which resembles the microenvironment in the lungs filled with air on one side and blood on the other. One of the novelties of this approach is the use of porous and flexible membranes that can stretch upon mechanical stimulation, mimicking the stretching of the lungs during inhalation (Fig B). They further showed that the lungs-on-chips showed organ-level responses to inflammation/bacterial infection, toxicity and absorption of nanoparticles, and cancer drug IL-2-induced pulmonary edema, similar to in vivo animal models. This new technology is promising because it can mimic how lung cells behave in both normal and diseased conditions; however, this is just one microenvironment out of many. Fortunately, the group has shown success in re-creating other organs on chips such as the intestines, kidney, bone marrow, and liver. Now, as Dr. Ingber posited at the end of his talk, we must await the day when all these chips are combined to form a single device of human-organs-on-chips.
Image Sources
Human Organs-on-Chips
Figure from Huh, et al. Science 2010.
All others were drawn/created by the author.
Works Discussed
Fogel. Contemp Clin Trials Commun, 2018. PMID: 30112460
Akhtar. Camb Q Healthc Ethics, 2015. PMID: 26364776
Huh, et al. Science, 2010. PMID: 20576885
Huh, et al. Science Translational Medicine, 2012. PMID: 23136042
Musah, et al. Nat Protoc, 2018. PMID: 29995874
Tovaglieri, et al. Microbiome, 2019. PMID: 30890187
Leave a comment