Reading time: 5 minutes
Shaye Hagler
The discovery of accelerators and brakes in our immune system in the early 1990s was fundamental to designing the cancer immunotherapy platforms we use today, which is why it won the 2018 Nobel Prize in Medicine or Physiology. Now, immune checkpoint 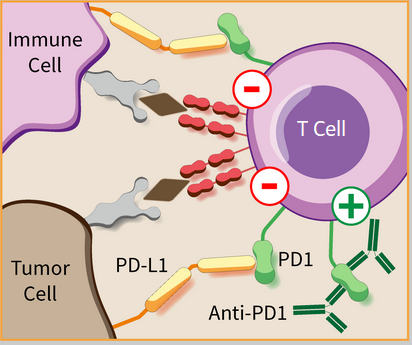 inhibitors like anti-PD-1 can work in tandem with surgery, radiation, and chemotherapy to help patients with aggressive tumors survive longer. Briefly, anti-PD-1-based therapy works by blocking a protein (PD-1) on the surface of T cells that, when bound to its counterpart (PD-L1), causes the T cell to shut down. In a healthy person, PD-1 tells the immune system not to overreact and attack healthy cells. However, tumor cells can hijack this process by making their own PD-L1 and thus avoiding confrontation with killer T cells. Blocking the interaction between a tumor cell’s PD-L1 and a T cell’s PD-1 enables your body’s own natural defenses to help defeat cancer.
inhibitors like anti-PD-1 can work in tandem with surgery, radiation, and chemotherapy to help patients with aggressive tumors survive longer. Briefly, anti-PD-1-based therapy works by blocking a protein (PD-1) on the surface of T cells that, when bound to its counterpart (PD-L1), causes the T cell to shut down. In a healthy person, PD-1 tells the immune system not to overreact and attack healthy cells. However, tumor cells can hijack this process by making their own PD-L1 and thus avoiding confrontation with killer T cells. Blocking the interaction between a tumor cell’s PD-L1 and a T cell’s PD-1 enables your body’s own natural defenses to help defeat cancer.
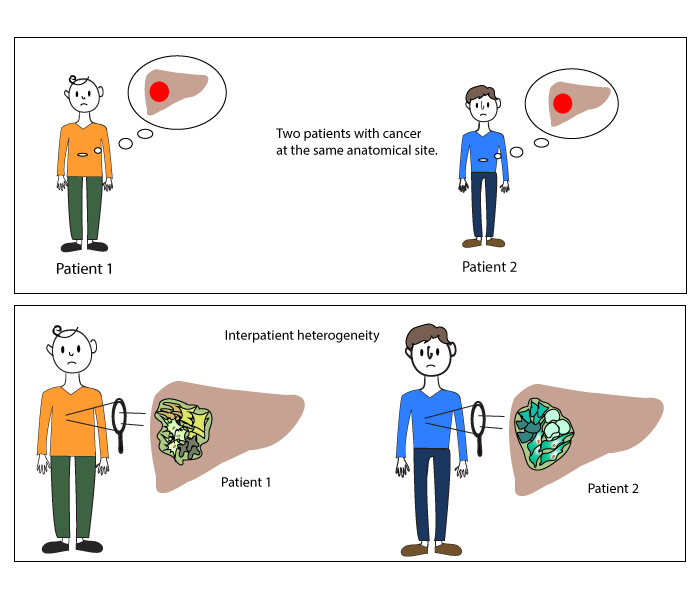
As always, however, there are limitations to this treatment approach. One of the main challenges is that not every patient responds to immune checkpoint inhibitors. Doctors try to avoid that by screening patient tumors for PD-L1 levels or other molecules involved in immune checkpoint inhibition. Unfortunately, there is more to treatment resistance than whether or not the interaction happens. Researchers at the Broad Institute of MIT and Harvard set out to identify methods to predict patient response to immunotherapy and discover strategies to improve response to therapy for patients who don’t respond well on their own. Using data about the different genes in different patient tumors allows scientists and doctors to recommend personalized treatment strategies, which is especially important for diseases like cancer that are both highly complex and require effective treatment quickly.
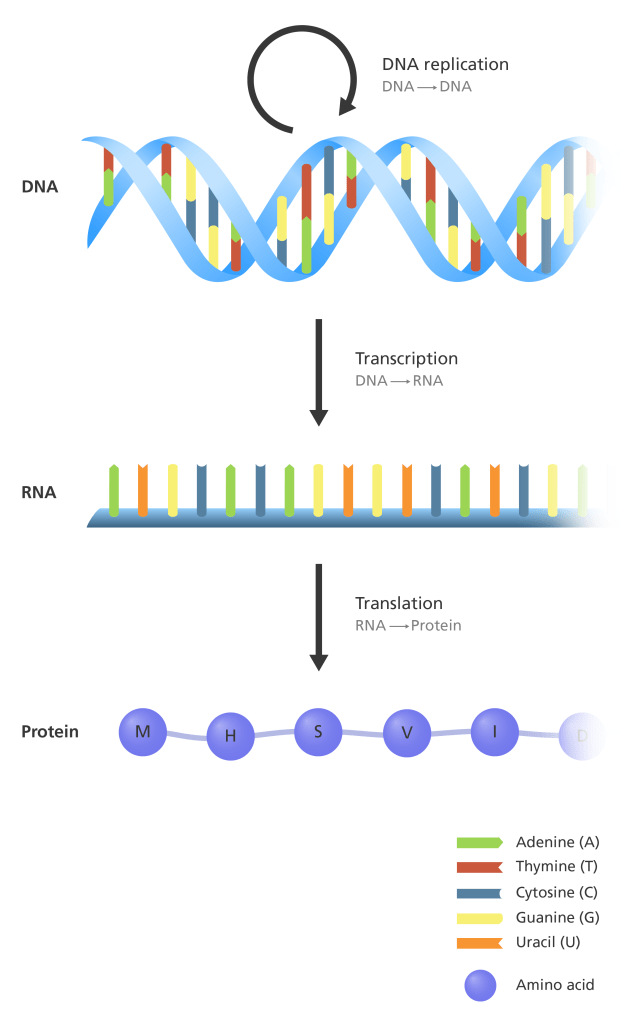
In order to understand more about how genes influence treatment response, the research team tackling this project used a method called RNA sequencing that reveals what types of and how much RNA exists in a given sample (in this case, they used patient tumors as their samples). RNA is the molecule that is made from genes and tells the cell what protein to build. Understanding what type of RNA exists is helpful to researchers as it can tell them what proteins actually get made; there are genes throughout your DNA that don’t ever get used! Furthermore, while the DNA stays the same, the RNA is constantly changing and reflects what’s actually happening inside each individual cell.
The research team at the Broad Institute looked at RNA in the tumors of different skin cancer (melanoma) patients before and after immune checkpoint therapy using two different methods: bulk sequencing and single cell sequencing. The difference between those methods might be clearer if we think of a tumor like a bowl of salad, and a pretty fancy one at that – with different kinds of lettuce and lots of toppings. If you want to find out what the bulk of the salad is made up of, you might dump out the bowl and mix it up evenly and then tally up everything in it. But that wouldn’t give you any information on where you’re more likely to find each topping and why that’s the case. That’s where single-cell sequencing comes in; you can figure out exactly where every crouton is in the salad and if it’s crispy or soggy. You can then use that information to predict where the croutons will fall in other salads and when they will be crispy or soggy. Except instead of croutons, this research team used this technology to look at T cells and which parts of tumors they could invade. What they found was a pattern of gene expression that could predict whether or not patients would respond to immune checkpoint therapy and do it better than the predictors we use now.
But they didn’t stop there. Next, they asked if there was something they could add to traditional immune checkpoint inhibitors to make them work on patients whose tumors don’t normally respond to that type of treatment. They tested 131 different drugs across 639 human cell types (!!) and discovered that a class of drugs called CDK4/6 inhibitors could fix the pattern of gene expression that keeps patients from responding well to therapy. In order to tell if that would make normally non-responding patients sensitive to immune checkpoint inhibitor therapies, they implanted resistant tumor cells in mice and then treated the mice with anti-PD-1 alone or anti-PD-1 plus a CDK4/6 inhibitor. The highest survival rates were associated with the mice who got the combination therapy, showing that drugs can be used to make immune checkpoint therapies work not just on normally responsive patients but also on patients who are predicted to be resistant to this strategy. Research like this is essential to advancing cancer therapy; there are plenty of drugs already out there that could help more patients if we can understand why they don’t work on certain populations. Expect to see cancer research move more and more toward a personalized medicine approach in the next couple of decades as a result of exciting data like these.
Work discussed:
Jerby-Arnon, L., Shah, P., Cuoco, M. S., Rodman, C., Su, M. J., Melms, J. C., … & Wang, S. (2018). A Cancer Cell Program Promotes T Cell Exclusion and Resistance to Checkpoint Blockade. Cell, 175(4), 984-997.
Image Credits
PD-1 immunotherapy. Pembrolizumab is an antibody that inhibits the abnormal interaction between the molecule PD-1 on immune cells and the molecule PD-L1 on tumor cells, allowing the immune cells to activate and attack tumors. Credit: CC BY-SA 3.0 – Guido4 (hegasy.de).