Reading time: 3 minutes
Rachel Cherney
If you have read other articles on Oncobites, you know that cancer is a collection of many diseases, and these diseases are complex. Cancer cells don’t live on their own; rather, they live among the normal tissue cells, immune cells, and bacteria in your body. Additionally, cancer cells can be different from one another, depending on whether they have gained mutations. This heterogeneity of the cancer cell environment makes studying cancer in laboratories particularly difficult.
In the case of drug studies, cancer cells are often grown in dishes where a drug of interest is applied, and scientists observe whether the cells are affected by the drug, and if so, how. However, many times only one cell type is used in the study, and this single cell type doesn’t reflect the complexity of the cancer cell environment. A drug that may have been effective on a single cell type may not have as much of an effect, or may have adverse side effects, in an actual patient.
How, then, are we supposed to adequately study cancer and possible treatments if simple models aren’t sufficient? Scientists are starting to turn to organoids to model tissue systems from which cancers can arise. Organoids are small, 3D organs that grow from stem cell culture. They are simpler than the organs in our own bodies, but more complex than a singular cell type in a dish. So far, organoid models have been developed for several organs, including the brain, lung, liver, and intestines.
A study by a group of researchers based mostly in the UK has looked into the benefits of using organoids for cancer studies. They demonstrated that not only could organoids model healthy organs, but they can also develop cancer to model diseased organs. By taking Primary Liver Cancer (PLC) tumors from patients, the researchers successfully grew liver organoids that replicated the cancer of the original patient. Using patient-derived cells means that, in the future, scientists and doctors could find personalized medicines for patients’ cancers.
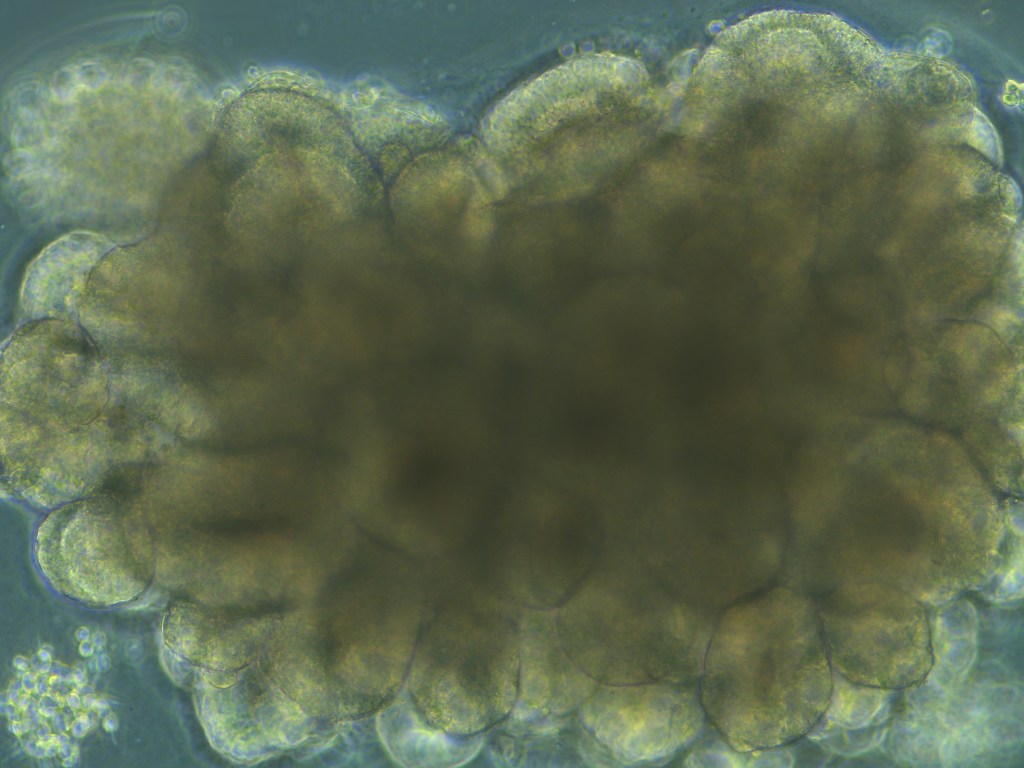
3D organoid
After developing patient cancer-derived organoids, the scientists wanted to see if cancer organoids could provide insight for novel PLC biomarkers. By comparing the expression of genes in healthy liver organoids and PLC organoids, they identified 11 genes with more expression in PCL than in healthy liver tissue, and four of these genes were associated with a worse cancer prognosis. Additionally, they performed drug sensitivity tests with 29 different drug compounds on multiple patient-derived PLC organoids. While many of the compounds had no effect on the cancer organoids, a compound targeting Erk1/2, proteins broadly involved in cell growth, stopped the function of Erk1/2 and prevented tumor growth. This is the first reported identification of an Erk1/2 inhibiting compound inhibiting PLC growth. While the Erk1/2 inhibiting compound would still need to go through clinical trials, the use of organoids shows promise in identifying novel drugs to treat cancer.
With continued scientific advances, it is now possible not only to grow miniature organs, but to grow organs from patient derived tumors. And it is only a matter of time before organoid models for other organs are developed. We now have an extremely useful tool for studying cancer in a complex environment and for determining which drugs are more likely to treat patient tumors before patients have their first treatment.
Edited by Jessica Desamero
Works Discussed:
Barbuzano, J. (2017, Nov. 7). “Organoids: A New Window into Disease, Development and Discovery.” Harvard Stem Cell Institute (HSCI). Retrieved from hsci.harvard.edu/organoids
Broutier, L., Mastrogiovanni, G., Verstegen, M. M., Francies, H. E., Gavarró, L. M., Bradshaw, C. R., … Huch, M. (2017). Human primary liver cancer-derived organoid cultures for disease modeling and drug screening. Nature medicine, 23(12), 1424–1435. doi:10.1038/nm.4438
Organoid Image Credit: https://www.flickr.com/photos/nihgov/48062903426/
Main Image Credit: https://www.needpix.com/photo/173917/brain-jar-conservation-mad-scientist-science-life-biology-experiment-thinking
Thanks for sharing this new innovation. It sounds like this could help groups have a more reliable go/no-go checkpoint for more time consuming and expensive downstream studies. And the patient derived aspect certainly adds an additional benefit.
LikeLike